Optical germ detection
Fast processes for bioburden detection are becoming increasingly important. Not least because of the ever closer internationally and globally entangled production chains and higher degrees of automation, contamination can spread very quickly very widely.
When it comes to germ detection, the first association is often hospitals, because there, of course, strict hygiene must be observed and compliance with the specifications must be checked.
Systemic prevention approaches (HACCP, GMP, etc.) are, however, also an integral part of everyday work in many other industries and are increasingly prescribed by standards. They play a major role, for example, in drug and food production.
The aim of preventive approaches is always to secure the production processes themselves through certain processes in such a way that infections or contamination with harmful substances or toxins are prevented before people fall ill, contaminated food is delivered or product recalls are started in order to contain the damage.
Hygiene processes play a central role, for example when cleaning tools and surfaces.
But how do you know whether cleaning a surface has successfully killed all germs? Many factors play a role here: the type and amount of cleaning agent, the type of surface, how the cleaning itself is carried out, the frequency of cleaning, etc. etc. And perhaps you are doing too much work and half the effort is enough to reliably sterilize a surface.
The result of the cleaning must therefore be checked. That is the job of germ detection methods.
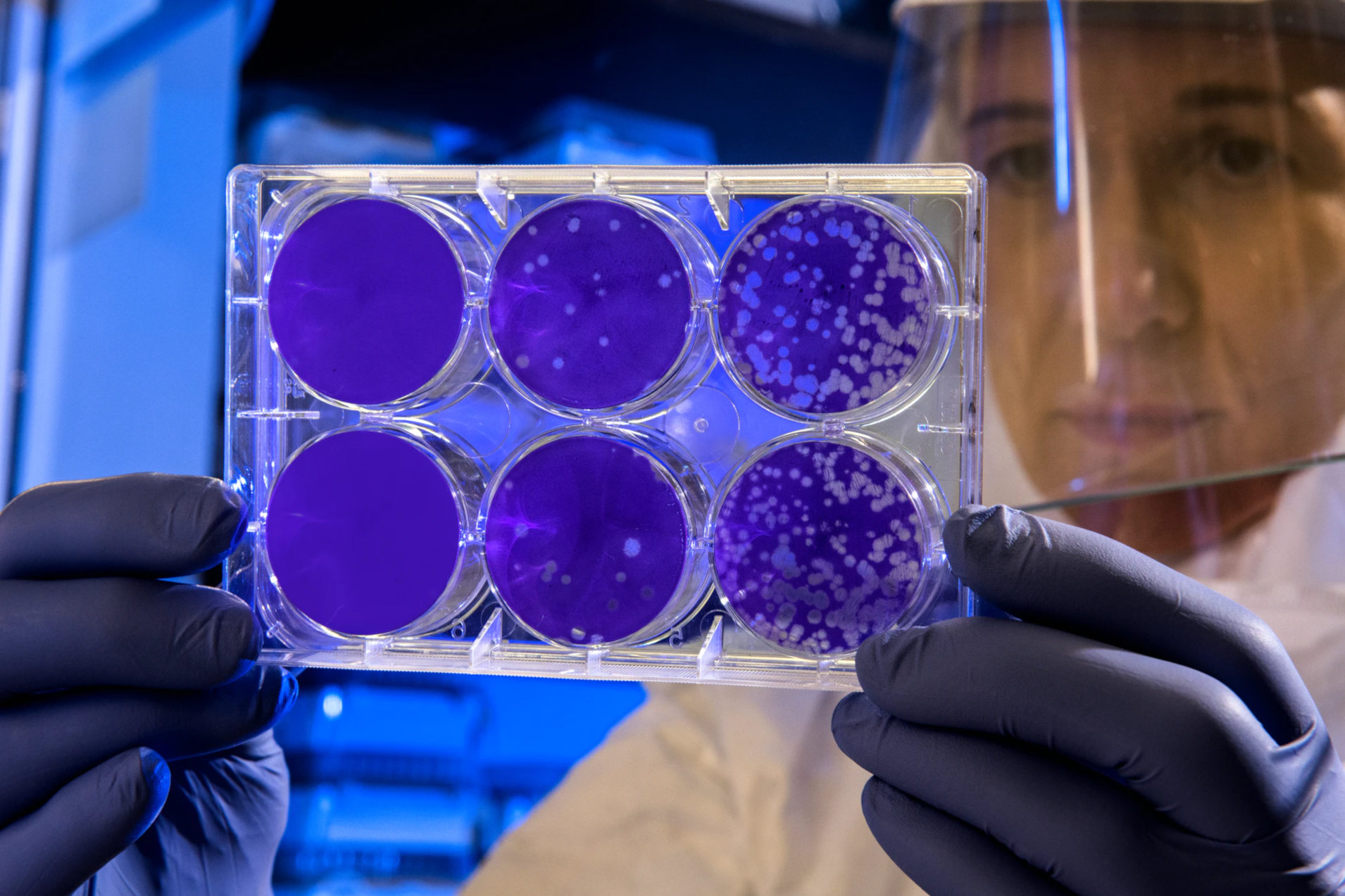
In this case, samples are traditionally taken from the surface, transferred to Petri dishes with nutrient solution and incubated for a few days, i.e. stimulated to grow in an environment that is as germ-friendly as possible. After a few days and many multiplication cycles, the germs have multiplied so much that the colonies are visible to the naked eye as different spots on the nutrient solution.
These spots, the Colony Forming Units (CFU), are then counted.
This process is robust and proven, but it is also very slow. For this reason, various rapid methods for identifying germs have been developed, which provide at least equivalent results in a significantly shorter time.
ATP bioluminescence
One of the most common rapid microbiological methods uses the same chemical reaction that makes fireflies glow.
In an ATP rapid test, a swab is used to take a sample from the surface to be tested and then put it in a cuvette (similar to a test tube). If there were germs in the sample, then there is now adenosine triphosphate (ATP) in the container, because ATP is contained as an energy carrier in the vast majority of cells.
A mixture of luciferase and luciferin is also filled into the cuvette. Together with the ATP, the luciferase-luciferin reaction occurs, during which the ATP is broken down. Excess energy is released during this process: For every ATP molecule a light quantum is emitted into the environment.
This is the source of the glow or luminescence. The more cells, i.e. ATP, was contained in the sample, the stronger the glow will be.
The result is still quite faint and, above all, we cannot distinguish the relative differences in intensity with the naked eye with sufficient accuracy. The evaluation of the bioluminescence is therefore mostly done with the help of machines.
Since different types of germs produce different levels of luminosity, it is unfortunately not easily possible to convert the luminosity exactly into the number of colony-forming units. In principle, however, it is possible if it can only be a certain germ whose ATP bioluminescence has been measured beforehand.
ATP rapid tests are easy to use, relatively cheap and the mobile devices are rarely larger than a pack of coffee, which is why this test form is very widespread.
DEFT (Direct Epifluorescent Filter Technique)
The DEFT method also works optically, but uses fluorescence, not luminescence.
A sample is taken again, diluted with a solution and then filtered through a special membrane. The solution also contains two different fluorescent markers, special dyes that only adhere to specific parts of a cell and glow at a specific wavelength when they are stimulated by other light.
DEFT uses a fluorescent marker to mark esterase, a substance that plays an important role in the metabolic activity of a cell. An active metabolism means that this cell is alive. This dye marks living cells.
A second dye binds to DNA. It is known that this is normally protected in the cell nucleus, so in order to reach the DNA the cell nucleus must be accessible, i.e. this cell is dead.
If the sample is then irradiated with exactly the light that causes these two dyes to fluoresce, the colony-forming units can then be counted using a microscope. Originally this was done manually, nowadays there are more and more devices where the counting is done by software.
The sample preparation is relatively complex and the laboratory equipment is not exactly cheap, which is why DEFT is mainly used for certain niches in which the very high precision of this technology is particularly advantageous.
Flow cytometry or solid phase cytometry
Flow cytometry makes use of the fact that many cell components show a very specific autofluorescence when irradiated with certain light frequencies, i.e. they display fluorescent properties by themselves without the addition of a special dye.
The detection of germs with the help of flow cytometry also begins with the removal of a sample. Here, too, the swab is first treated with a solution in order to get the germs from the swab into the liquid.
This liquid is now sent through a very narrow channel so that practically only individual cells can pass through the narrowest point, one at a time.
At this point, several laser beams are used to irradiate very specific wavelengths from different directions and the result is measured with the help of photodiodes. The whole thing happens very quickly, more than 1000 cells per second can be checked without any problems.
Due to the characteristic fluorescence responses and intensities, many different cells can not only be counted, but even identified in this way.
Flow cytometry is most commonly used for samples that are already liquids, such as drinking water or blood samples. Fluorescent dyes can also be used here in order to be able to count dead cells separately, similar to DEFT.
PCR (polymerase chain reaction)
Although only employing optical technology in the last step, due to the corona pandemic, PCR tests are a method often mentioned nowadays.
In PCR tests, genetic information is detected quite directly. To do this, the genetic material in the sample has to be duplicated. In living cells, DNA is present in the familiar double helix form. In the case of natural cell division, this double chain is chemically separated and a new counterpart is produced in each case, so that at the end of the process two identical chains exist.
The PCR process mimics these processes in the laboratory. Here however the separation takes place with the help of high temperatures: If the DNA chain is heated to approx. 95 °C, it will split open by itself.
A polymerase is also added to the sample, which enables the DNA to replicate. The necessary DNA building blocks, the deoxyribonucleoside triphosphates, are also added.
It is currently not possible to replicate the entire DNA in this way, only certain sections. However, this is not a disadvantage in the case of germ detection, because here you are usually interested in very short sections. For example, there is a certain short section that all bacteria contain and that is therefore often used for germ detection.
The beginning and end of the desired section are marked with so-called primers, which are also mixed in. In the first cycle after the separation, one primer binds, in the next also the second, so that the PCR product is only present in the correct length at this point in time.
The cycles of heating and subsequent replication thus lead to the duplication of the section of interest. It is therefore repeated many times depending on the original concentration.
In the case of a contaminated sample, there would now be a sufficient number of PCR products to enable reliable detection.
For this purpose, the sample is placed in a special, electrically conductive gel. When a voltage is applied, the DNA strands are sorted according to their length. At the same time, a so-called DNA ladder is added as a measuring stick, which contains different strands of known length and thus enables a direct comparison of the sample with the reference lengths. So far, so good, only the material now has to be made visible. Here, again, fluorescent markers that bind to DNA come into play. These are illuminated with the appropriate wavelength and then emit at a different, known wavelength, which is mostly in the visible range, but can also be located in the long-wave UV range.
PCR tests can detect very low levels of germs, but take several hours and require the appropriate laboratory equipment.
Outlook
The rapid microbiological methods shown can in some cases detect very small amounts of colony-forming units - but the precision is bought at the price of longer test periods and higher costs. This makes them very suitable for random checks. For continuous measurements, which can be carried out regularly as an element of a HAACP or GMP process to determine bioburden, additional techniques would be desirable that could provide sufficiently precise results directly in the field within a few minutes.



